|
VIPVertically Integrated Project: Undergraduate Research Team
|
|
Virtual Journal ClubCOVID-19 has shown us that some things, like journal clubs, can be effectively done on-line. This opens up the possibility to include participants from different universities or even companies, as we are constrained only by time-zone differences. Both trainees and PIs are encouraged to join, but the goal is to make this a trainee-based discussion with minimal PI input except to clarify and steer the discussions. |

|
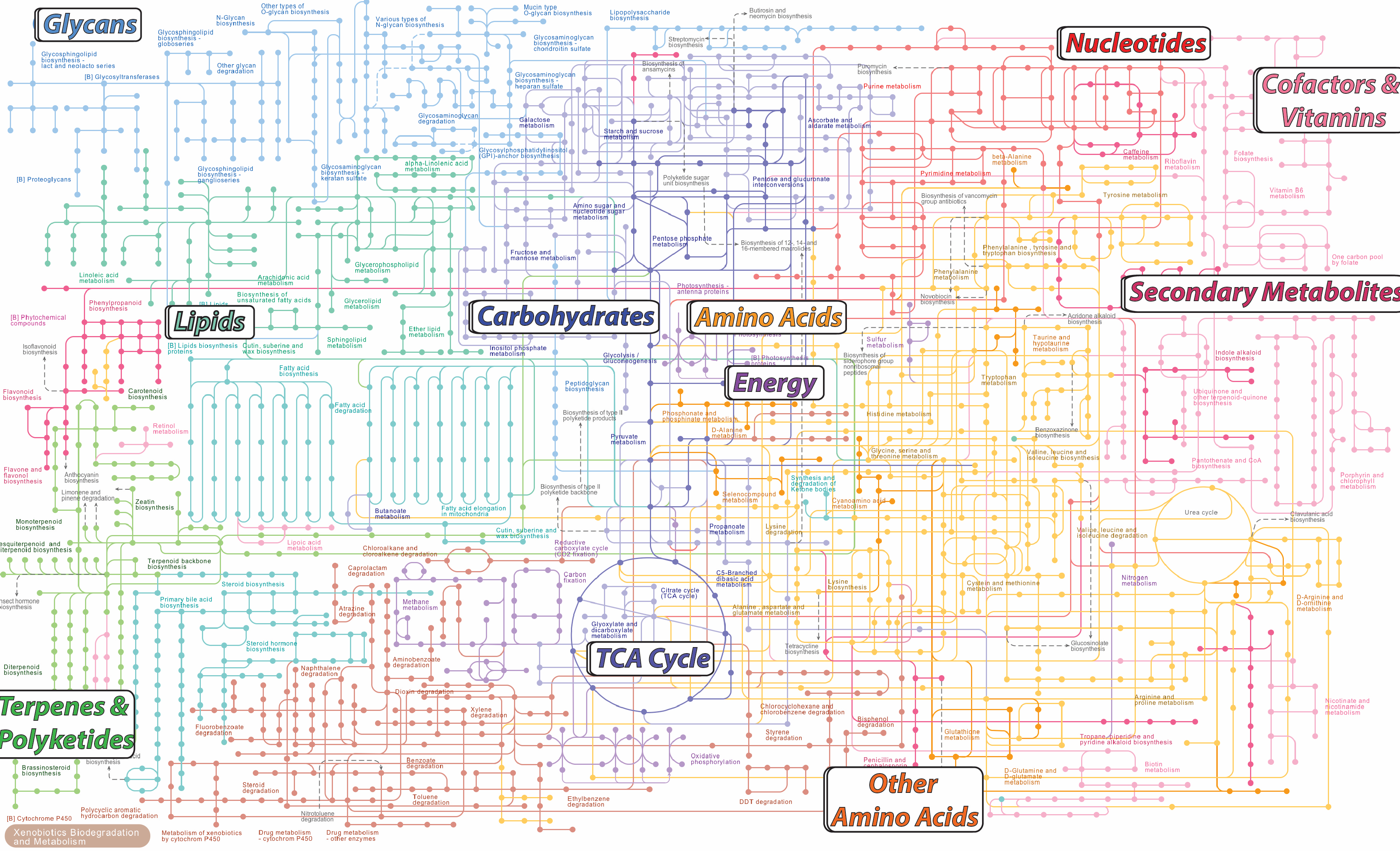
Introduction to BioinformaticsThis class is offered every fall semester, and taught with Jim Leebens-Mack and Jonathan Arnold. It is for advanced undergraduates or graduate students. The overall class focuses on systems biology, with introductions to omics (metabolomics, genomics, and RNAseq), an overview and introduction to trees for analysis of phylogentic and omics data, and an introduction to programming in MATLAB. The 3 instructors attend and participate in most of the classes, so it is not a traditional team-taught class. |

|
Lecture notes in NMRI have compiled a set of 9 lectures that provide an overview of NMR. I hope to update these soon. |